Our work
Our research explores how cells react to/leverage small molecules, work that takes us through a diverse range of experimental strategies and approaches.
Antimicrobial resistance
Bacteria produce small molecules to interact with their environment, including those used to fend off their enemies. These molecules have been repurposed as antibiotics, revolutionizing healthcare. Unfortunately, bacteria can readily develop antibiotic resistance, and the spread of resistant bacteria threatens to undo many of our advancements in modern medicine.
We developed an easy and scalable evolutionary platform, Soft Agar Gradient Evolution, or SAGE (1). Through SAGE we can generate resistant bacteria in as little as sixteen hours, allowing us to study how resistance evolves and identify ways by which we can slow the pace of evolution.
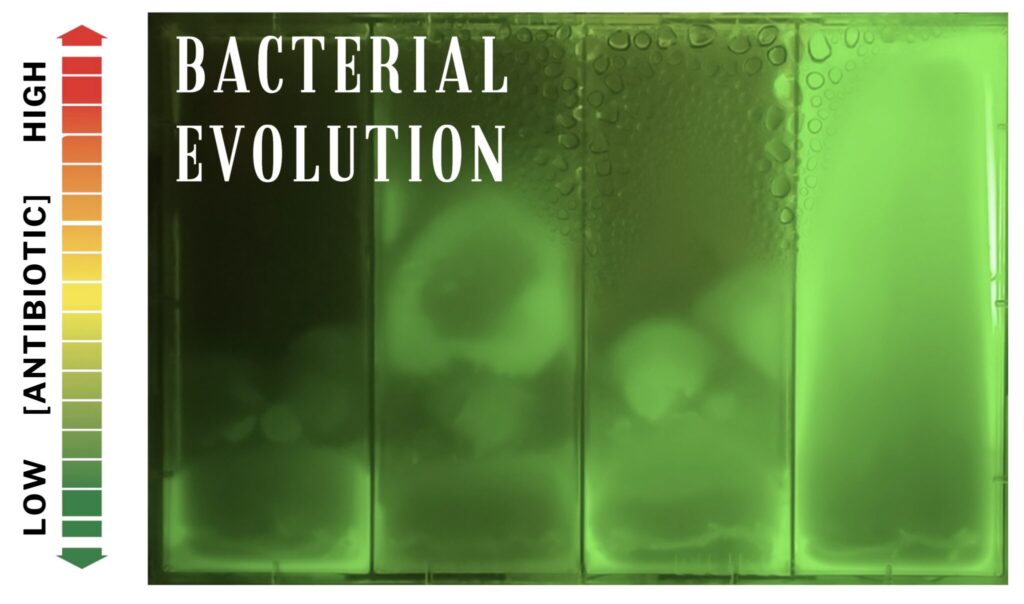
SAGE plate
We also believe there is an urgent need to include antimicrobial resistance evolution in the evaluation of potential new antibiotics. We are currently using SAGE to predict resistance mechanisms and develop strategies to limit the spread of antimicrobial resistance before it appears in the clinic.
Bacteria often develop resistance at a cost – a collateral effect that we may exploit to develop new antibiotics. However, bacteria’s diverse resistance mechanisms make predicting these effects challenging. We work to identify collateral effects of resistance that are predictable, and to then leverage them to build antibiotic therapies that are more resilient against resistance evolution. (2, 3, 4)
Chemical ecology
We found that bacteria use the small molecule geosmin as a signal to deter predators, marking the first known instance of a bacterial warning chemical. Geosmin influences the behavior of C. elegans, acting as a chemical warning that indicates toxic, unpalatable bacteria, thereby reducing predation and benefiting both the bacteria and the predators. This is the first aposematic signal to be reported in bacteria, and one of the first chemicals to act principally as an aposematic signal. (10, 11)
Compound discovery
We collaborate with national and international laboratories to test the efficacy of novel antibiotic therapies and fundamental studies on mechanism of antimicrobials, and have dedicated projects aimed at identifying potential new antibiotics from both bacteria and chemical libraries. (5, 6, 7, 8, 9)
Small molecules like adjuvants can restore the efficacy of antibiotics against resistant bacteria. Repurposing a drug library, we identified perforone, which enhances the action of polymyxin B in polymyxin-resistant strains of E. coli. (12)
Additionally, we use innovative techniques to upregulate biosynthetic gene clusters (BGS) to discover cryptic bioactive compounds. We are studying how bacteria produce small molecules in response to stress induced by the presence of other bacteria in the culture or a conjugated plasmid. We are identifying specific compounds produced across different species, with the potential to discover new antibiotics, and investigating which conditions trigger the strongest stress responses and natural product output.
Bioremediation of heavy metals
We harness our expertise on microbial evolution to adapt bacterial strains for bioremediation of mine waste. With help of the Lallemand Graduate Fellowship in Bioprocessing, we are attempting to create bacteria better able to remove heavy metals from tailings ponds, detoxifying the environment and serving as a valuable resource for critical minerals.

